All posts by Chris Wolverton
Fruit Development
- Fruit development
- Patterning
- Carpel development requires AG and SEP genes (MADS box TFs)
- Genes only in flowering plants, common to all
- Dry fruits need to have a means of opening, which is
regulated
- Carbohydrate import, storage as starch
- Polyphenolic synthesis
- Associated with pigment
- Also associated with lignification of dry fruit carpel wall
- Patterning
- Fruit ripening
- Starch conversion to simple sugars
- Wall breakdown
- Galactosidases, glucosidases
- Ethylene signaling
- Receptor family has five members of two types
- ETR1 and ERS1 form one type
- Ethylene binding and dimerization at N terminal
- His kinase domain at C terminal
- Like in bacterial two component signaling systems
- ETR2, ERS2, and EIN4 lack His kinase domain, have
different kinase domain
- ETR1 and ERS1 form one type
- Signaling pathway
- CTR1 is a negative regulator
- Mutant shows constitutive triple response
Radial swelling, hypocotyl inhibition, hook
opening - Encodes product homologous to MAPKKK proteins
- Mutant shows constitutive triple response
- CTR1 is a negative regulator
- Receptor family has five members of two types
Pollination
- Pollination
- self vs non-self recognition
- Self incompatibility ensures cross-fertilization
- S locus encodes components of recognition
- SRK a kinase expressed in stigma PM
- Found in papillae cells, interacts with SCR
- SCR expressed in pollen
- Peptide secreted by microspore and tapetal cells
- Issues of sporophyte or gametophyte origin
- SRK a kinase expressed in stigma PM
- Self rejection pathway
- Pollen complex with SRK endocytosed and degraded
- Same papilla can detect self simultaneously with cross
- pollen tube elongation
- Tip growth
- Ca++ gradient
- Actin necessary
- Water relations
- Water uptake from surrounding tract
- Tip growth
- egg cell signals
- self vs non-self recognition
Floral Organ Identity
As long as a shoot apical meristem has a vegetative identity, it continues to produce new leaf primordia that emerge in a particular pattern from the apical dome. When the repression of the floral meristem identity factors AP1 and LFY is released by the induction of flowering, the meristem switches to a program of producing the floral organs: sepals, petals, stamens, and carpels. These organs form in distinct bands, or whorls, that encircle the meristem, which leads to the radial symmetry found in most flowers. Even in flowers showing bilateral symmetry, the same organs are formed in roughly the same whorls, but the organs are often fused in distinctive ways that lead to novel flower shapes. The central question, then, is how does a meristem control the placement of the proper organ in the proper location — how are the floral organs specified?
ABC Model of Floral Organ Formation
Using a genetic approach to answer the question of floral organ identity, researchers have isolated mutants in floral organ formation that show specific defects in the formation of one or more organ. Intriguingly, when an organ went missing, it was often “replaced” by a different organ. For example, in one flowering mutant, the plant failed to form stamens and carpels, and instead replaced them with petals and sepals, respectively. This kind of mutation, where an organ is replaced with another, is known as a homeotic mutation: the replacement of one organ with a different one.
Screening for flower organ mutants eventually uncovered many genes involved in the specification of organ identity, and the genes were often named based on the phenotype of the mutant. The APETALA mutants, AP1, AP2, and AP3, all lacked petals. The PISTILATA (PI) mutant produced excess pistils in place of stamens. Finally, the AGAMOUS (AG) mutant lacked both stamens and carpels, the gamete-bearing organs. As more and more of these mutants were isolated and characterized, a pattern began to emerge that would become known as the ABC model.
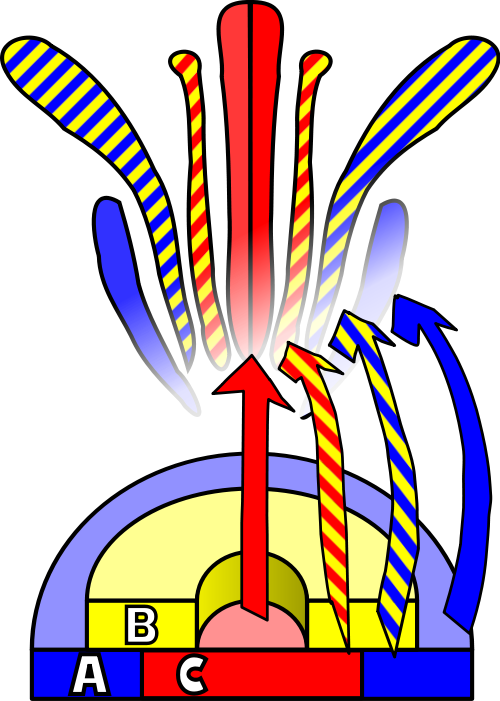
To explain the homeotic effects, it was postulated that class ‘A’ genes and class ‘C’ genes mutually exclude each other. In cells where the ‘A’ genes are expressed, there can be no ‘C’ gene expression. This phenomenon is common in development, with ‘A’ and ‘C’ known as cadastral genes, setting up a firm boundary between them. Returning to the ‘B’ class, these genes are expressed in an overlapping fashion atop both ‘A’ and ‘C’ regions such that, when ‘B’ mixes with ‘A’, the result is a petal and when ‘B’ mixes with ‘C’, the result is a stamen.
If we return for a moment to the mutant phenotypes, we see that this model explains the main observations of each mutant. For example, the loss of AP1 by mutation results in the pattern carpel-stamen-stamen-carpel. The domain of ‘C’ expression expands into ‘A’ territory, just as would be predicted by loss of ‘A’ activity, resulting in the specification of organs associated with C alone (carpel) or B plus C (stamen). Likewise, agamous mutants show the pattern sepal-petal-petal-sepal, just as predicted by the loss of ‘C’ activity and explained by the encroachment of A activity to take its place. The organs specified are those of A alone (sepal) or A plus B (petal). Finally, in the case of the pistilata mutant, the pattern observed was sepal-sepal-carpel-carpel, just as predicted by the loss of ‘B’ activity.
Since its initial proposal, the ABC model has been enhanced and refined. A fourth class, known as ‘E’, has been added to recognize the contribution of the SEPALLATA genes 1-4. These act as co-regulators in all whorls of the floral meristem, and the loss of all SEP genes results in the formation of only sepals in all organs, thus returning the flower to a collection of leaf-like organs.
The cloning and molecular characterization of the ABC genes revealed them to mostly encode transcription factors. The common thread that ties them together is the nature of the DNA element to which they bind, known as the MADS box. The MADS box is a DNA regulatory region characterized by the sequence CC[A/T]6 GG, and is present in the targets of ABC gene activity. Since their initial discovery in the model plant Arabidopsis, homologs of these MADS-box transcription factors have been identified not only in the flowering plants, but across all eukaryotes.
Transition to Flowering
One of the most significant developmental changes a plant experiences is the switch to reproductive growth from vegetative growth. This transition requires a wholesale change in the identity of the shoot apical meristem, and for most meristems it also means a shift from indeterminate to determinate growth, implying that any new leaf growth must come from another meristem. Given the biological significance of this transition and the horticultural interest in understanding and controlling the timing of flowering, the events that lead to flowering have been intensely studied.
Photoperiod
As with so many other ‘decisions’ made by plants, the inputs that regulate the transition to flowering are made up mostly of cues from the environment. For most plants, the strongest influence over the transition to flowering is day length, and plants can be sorted into 3 classes based on the photoperiod required to induce flowering. Short-day plants transition to flowering when day length begins shortening and reaches a duration less than the critical period for that species. In the northern hemisphere, these are plants that flower after June 21, the longest day of the year. In contrast, long-day photoperiodic plants begin their transition to flowering as day length increases past a certain minimum length. In the northern hemisphere, these are plants that flower in spring and early summer, as day length is waxing. Finally, many plants are day-neutral and will transition to flowering regardless of light cues. Plants that have strict photoperiod requirements for flowering are known as obligate short- or long-day flowering plants, while those that will eventually flower regardless of day length but a particular photoperiod promotes flowering are known as facultative short- or long-day flowering plants.
There is a certain irony in all this talk of day length in promoting flowering, which is that the plant is not actually responsive to the day length at all, but rather to the length of the dark period. In order for obligate short-day flowering plants to begin flowering, they must experience a long, uninterrupted period of darkness. Likewise, obligate long-day flowering plants must experience a shorteningn period of darkness. The reason plants sense the duration of darkness lies in the biophysics of phytochrome, the red/far-red photoreversible photoreceptor. The pool of phytochrome molecules becomes activated by exposure to red light, such as that experienced by a plant during daytime. This pool of molecules slowly reverts back to the inactive red-absorbing form during the dark period, hence, the longer the duration of darkness, the greater the proportion of total phytochrome is inactive. For obligate short-day (or long night) flowering plants, if this dark period is interrupted by even a brief pulse of intense light, that is enough to disrupt the balance of phytochrome signaling that leads to floral transition.
Other cues
A number of other signals besides photoperiod can influence the transition to flowering in some plants. Some plants flower after a given number of days since germination, regardless of the photoperiod. Related to this, some plants transition to flowering after attaining a minimum size — for example, many trees must be 5 or more years old before flowering, probably due to the high resource demands associated with seed and fruit production.
Temperature is another particularly important cue for some plants in the transition to flowering. Many plants require the exposure of the meristem to cold temperatures in order to complete the transition to flowering meristem identity. This process of cold treatment is known as vernalization because of its association with growing through the winter months. The cold treatment has been shown to modulate certain epigenetic factors associated with gene expression, such as DNA methylation. Some plants that have this requirement undergo a two year life cycle with the exposure to cold in the intervening winter. These plants are known as biennial plants, flowering in their second year of growth and typically dying thereafter.
Molecular Signaling
In order to begin making flowers, shoot apical meristems must undergo a change in identity from vegetative to floral (or inflorescence) meristems. This change in meristem identity is influenced by numerous environmental cues, but how do these cues affect the identity of the meristem? Where do the signals come from that result in flower formation, and how do they form?
Through a series of clever experiments starting in the 1920’s, plant biologists have shown that, although the meristem is the site of floral transition, the signals that convert the meristem from leaf-making to flower-making come from the leaves. One of the key experiments to demonstrate this involved grafting a leaf from a plant induced to flower onto a plant that had not yet been induced, resulting in the induction of flowering. For many decades researchers attempted to isolate the diffusible substance causing flowering, but were unsuccessful. Around this same time the plant hormones auxin and gibberellin had been succesfully isolated and characterized, leading many physiologists to search for a similar small organic molecule as the “flowering hormone,” or florigen.
Recent work has revealed that the mRNA and/or protein product of FLOWERING LOCUS T (FT) acts as a positive regulator of flowering. Its expression is upregulated in response to photoperiodic induction in the leaves, and it trafficks through the phloem to the shoot apical meristem. FT expression in leaves is mediated by the expression of another gene, CONSTANS, which encodes a transcription factor that is expressed as an output of the photoperiodic pathway. The movement of FT protein and/or mRNA from the leaves to the meristem requires interaction with a partner called FTIP, or FT interacting protein.
Upon reaching in the meristem, FT interacts with the transcription factor FLOWERING LOCUS D (FD), and together they activate expression of genes that change the identity of the meristem to that of a flower. One example target of the FT-FD dimer is APETALA1 (AP1), a meristem identity gene; another example target is SOC1, which promotes the expression of another meristem identity gene called LEAFY (LFY). Both AP1 and LFY expression are repressed during vegetative growth, and this repression is removed upon arrival of FT in the meristem.
Reproduction
How do plants reproduce?
- Transition to Flowering: How does a plant switch from making leaves to flowers?
- Floral Organ Identity: How are flowers made?
- Pollination: How is the sperm cell delivered to the egg cell in plants?
- Fruit Development: How do the parts of the flower develop into a fruit?
Defense Signaling
One of two major response pathways is initiated when plants are attacked by microbial pathogens. One major difference difference between these two pathways is the specificity with which the plant identifies the pathogen. In one pathway, the plant recognizes general signals common to a broad array of pathogens. These signals are known as pathogen-associated molecular patterns (PAMPs), thus this pathway is called PAMP-triggered immunity (PTI). In the second kind of signaling pathway, the plant responds to highly specific molecules secreted by the pathogen. These molecules are called effectors, hence this pathway is known as effector-triggered immunity (ETI).
PAMP-triggered immunity (PTI)
The hallmark of PTI is the ability of the plant to detect the molecular signature of pathogens. These signatures may take many different forms, including peptides derived from bacterial flagellum proteins, chitin (which makes up the cell wall of fungi), double-stranded RNA, and certain sequences of DNA common to microbes. Each of these presents a specific molecular pattern recognizable by the plant as indicating the presence of a potentially pathogenic organism.
These molecular signatures are detected by the cell through a class of proteins known as pattern recognition receptors (PRRs) that are specific for certain molecules. For example, the flagellum peptide known as flg22 is perceived by a PRR called FLS2. It is important to note that the flg22 peptide is a common motif found in all bacterial flagella as they are hydrolyzed. Upon perceiving flg22, FLS2 interacts with and phosphorylates the first member in a kinase cascade called MEKK1. MEKK1 phosphorylates one or more of several kinases (MEK4/5), which target several other kinases (MPK3/6). This series of events is known as a mitogen-activated protein kinase cascade, a signal transduction pathway common across all organisms. This kind of signaling system is modular and achieves the amplification of a single signal. The end result of all of this kinase activity is the activation of several transcription factors (WRKY22/29) that control defense-regulated genes. One example of these genes is those involved in synthesis of salicylic acid and jasmonic acid. There is also evidence that the kinase cascade can modify chromatin structure through acetylation and methylation, both of which can have a strong effect on genome-wide gene expression. All of this signaling has the overall result of “priming” the plant defense response.
Effector-triggered immunity (ETI)
The fundamental difference between PTI and ETI is the degree of specificity, with ETI representing a highly-specific, gene-for-gene defense response. The high specificity makes this pathway less durable than PTI and more targeted against an individual pathogen.
The ETI response is initiated by the secretion of effector molecules by the secretory system of bacterial cells. These effector molecules, called Avr (avirulence) proteins, are detected by the plant through recognitional resistance genes, or R-genes. The R genes encode Ser/Thr protein kinases, which interact with partners called nucleotide binding leucine-rich repeat proteins (NB-LRRs). This signaling complex also touches of a kinase cascade, but the overall result is the activation of the hypersensitive response and rapid cell death around the site of infection.
The avirulence (Avr) proteins represent an interesting evolutionary riddle: how is it adaptive for the bacteria to secrete a molecule that signals its presence to the plant? In at least one case, the answer seems to be related to the function of the Avr protein. AvrAC, secreted by a strain of Xanthomonas, acts as a uridyl transferase that modifies a component of the signal cascade such that its phosphorylation site is masked. Unable to be phosphorylated, the kinase cascade is interrupted and the plant fails to activate the hypersensitive response.
Defense Overview
Plants are under constant attack in the environment, with the attackers trying to breach the perimeter by chewing, ripping, dissolving, penetrating, or tearing open the organs, tissues, and cells making up the plant body. These attackers include everything from acellular viruses and single-celled bacteria to multicellular mammals. Plants have evolved defense systems to address this wide variety of potential attackers. Those systems include physical barriers and anatomical structures to prevent the penetration of an attacker, general chemical defenses that act as a deterrent against attack, and highly specific immune-type responses that recognize and protect against familiar pathogens.
Kinds of Pathogens
In addition to the multicellular attackers such as nematodes and insect larvae, single-celled microbes are a constant threat to a plant’s health. This class of attacker falls into one of three possible categories depending on the mode of action employed by the pathogen. On one end of the spectrum, some microbes seek to destroy the plant tissue and absorb as much of the remaining organic matter as quickly as possible. This category of pathogen is known as the necrotropic pathogens due to the fact that these microbes feed on dead tissue. On the other end of the spectrum, some microbes attempt to penetrate the host plant’s defenses and soak up as many organic resources as possible without the host mounting a defense response. These are biotrophic pathogens, attempting to evade host detection and keep the host tissue alive because it benefits them the most to do so. These are the microbes that often induce tumor formation in host organs by transferring genes into the host cell that encode hormone biosynthesis enzymes, resulting in increased cell division. These pathogens are the equivalent of a squatter taking up residence in a home and building and addition onto the kitchen so they can get more to eat. In between these extremes are the hemibiotropic pathogens, which keep the host tissue alive for a time after invasion, but eventually kill it and take its resources.
Structural Defenses
The first line of defense plants present to an invading pathogen is the thick cuticle found on the surface of the epidermis throughout the aerial tissues of the plant. The cuticle is made up mostly of waxes and fatty acid esters known as cutins that are secreted by the epidermal cells. In addition to making the surface more water-tight, these waxes and cross-linked esters deter some smaller animal herbivores and many microbes.
While the cuticle forms an effective barrier on leaves and stems during primary growth, stems undergoing secondary growth secrete a different kind of coating for protection. These organs produce a layer of cells that contribute to an increase in girth of the stem by dividing in a ring around the stem parallel to the surface, forming cork. These cork cells, which make up part of the bark of stems, synthesize a waterproof substance called suberin, which is secreted into the apoplast and permeates the cell wall of cork cells. Once suberin is manufactured and secreted, these cells can no longer take up water and die. Suberin is a polyphenolic substance and is not only hydrophobic but also has some antimicrobial properties as well as providing a physical barrier.
Trichomes, or hair cells, provide a physical impediment on the surface of many stems and leaves that deters the feeding of many insects. Not only do trichomes offer a physical challenge, they are often filled with secondary compounds that are irritants or toxins to feeding insects or other animals. Cooking herbs are an abundant example, often having leaves covered in terpene-bearing glandular trichomes that burst when contacted. Another example is poison ivy (Toxicodendron radicans), which secretes an irritating oil from trichomes called urushiol.
If an attacker manages to make it through cuticle covering the surface of the plant, perhaps through a wound opening, it will still find one last line of defense: the tough, rigid, lignified cell wall. Made up of cellulose and other structural polysaccharides impregnated with the polyphenol lignin, the cell wall presents a formidable barrier for microbial attackers like bacteria and fungi. Some species of fungi form elaborate multicellular structures with their hyphae (called an appresorium) that can puncture the highly-pressurized cell wall, while other fungi and bacteria secrete cellulose-degrading enzymes that help to digest the wall away.
Defense Responses
Once the structural defenses have been compromised, the plant begins to mount one or several simultaneous defense responses leading to only a few outcomes. In many cases, the results of an attack include the increased biosynthesis of secondary compounds through the induced expression of genes encoding enzymes in terpene or phenolic compound synthesis. These molecules are always being synthesized by the cell at a baseline rate, but that rate increases when an attack is detected.
Another outcome of infection can be the initiation of programmed cell death at or near the site of infection. When the plant dectects certain features of a pathogen or herbivore, these can trigger those cells to cut themselves off from the rest of the plant to try to contain the infection. If you inspect a leaf, it is not uncommon to find brown spots scattered throughout the leaf. These spots are a common indicator of such cell death activity.
Pathogen and herbivore attack can also induce the plant to produce signal molecules that communicate an infection throughout the rest of the plant body. Salicylic acid and jasmonic acid are two such hormones that serve to prime the immune response of distant organs in advance that a pathogen or herbivore has attacked.
Secondary Compounds
Plants synthesize a wide range of metabolites that are not known to be essential for their survival. Because they are not a part of primary metabolism, they are often lumped together in a category of metabolites called secondary compounds. This is an entirely artificial categorization given that there is no unifying characteristic of the entire group. Many of these compounds are involved in defense responses, but others produce pigments involved in pollinator attraction, and still others make up important photoprotective pigments in leaves that guard against damage due to high light conditions. Most of the secondary compounds are strongly oxidizing, and thus are sequestered in the vacuole after production to protect the proteins of the cytoplasm from damage.
While the structures and functions of these molecules are diverse, they can be grouped into 3 major classes based on common structure and biosynthetic pathways. The classes include the phenolic compounds, terpenes, and nitrogen-containing compounds. No single plant species makes a full complement of all secondary compounds, but each makes a select subset of them.
Phenolic compounds
Phenolic compounds are united by having a phenol group — a phenyl ring with a hydroxyl group. Such structures are assembled through the shikimic acid pathway, which branches off of primary metabolism through phosphoenolpyruvate in the respiration pathway and leads to the synthesis of the aromatic amino acids phenylalanine, tyrosine, and tryptophan. Phenylalanine serves as the entry substrate to form the phenolic secondary compounds, with the first committed step being the deamination of phenylalanine by the enzyme phenylalanine ammonia lyase (PAL). The simplest resulting molecules are the simple phenolics, phenylpropanoids, and the precursors to lignin. When these simpler phenolics condense with a second phenyl ring, they form flavonoids, isoflavonoids, and flavonols.
One example of a common flavonoid is the family of pigment known as anthocyanins, which contribute the reds, purples, and blues to flower pigments. The different colors of anthocyanin pigments are the result of varying degrees of hydroxylation on one of the 2 rings of the flavonoid. Many other examples of flavonoids are encountered every day in the foods humans eat, specifically from the herbs and spices added to food for flavoring and spice. Flavonoids are the primary flavor constituents in such spices as cinnamon, ginger, cloves, nutmeg, coffee, and vanilla. These flavonoids probably serve as a deterrent to herbivorous insects and larger animals, and many also show antimicrobial properties.
Terpenes
Terpenes are long chain hydrocarbon molecules having 10-40 carbons. Most are toxic in some way to animals and so are deterrents to herbivory. In addition to these defense compounds, the plant hormones GA and ABA are examples of terpenes, as are the carotenoids, yellow-orange pigments involved in photosynthesis. Like the phenolic compounds, the terpenes are synthesized by a pathway that branches from respiration through either acetyl CoA or glyceraldehyde 3-phosphate and pyruvate. The results of either pathway are 5-carbon units that can be joined together in a variety of ways to produce the long chain structures associated with this class. As a consequence of this biosynthetic process, the terpenes always occur in multiples of 5 carbons (10, 15, 20, etc).
The monoterpenes are the smallest of the terpenes, composed of 10 carbons, and are often volatile essential oils found on the surface of leaves and stems. The familiar scent of pine, for example, is produced by α-pinene and β-pinene. Another familiar monoterpene is limonene, responsible for the lemon scent. In the case of both pine and citrus, these essential oils are found in a sub-epidermal duct or ‘oil gland’ where they are maintained until the tissue is disrupted. Most of the flavor compounds associated with the mint (Lamiaceae) family, including basil, oregano, mint, and thyme, are produced by monoterpene essential oils. In these examples, the oils are found in modified trichomes atop the surface of the leaf, where they are easily ruptured and released as a potential herbivore merely contacts the tissue.
Another class of terpene is the diterpenes (20 carbons) that are represented by pine resins, found in ducts throughout the stems of most pine species. Another important diterpene is the anti-cancer drug taxol, found in the bark of the Pacific Yew tree. This complex diterpene binds to microtubules and stabilizes them, resulting in an arrest of mitosis. The triterpenes (30 carbons) are large, complex structures that undergo ring condensation to form steroidal compounds and cardiac glycosides, so named because of their effects on vertebrate heart function. Several of these compounds are used to treat heart conditions, including digoxin, from the foxglove plant (Digitalis sp.) and oleandrin, from the oleander plant (Nerium oleander). Finally, the tetraterpenes (40 carbons) include latex and chicle.
N-containing compounds
Whereas representative molecules of both phenolics and terpenes are found in all plants, the nitrogenous compounds are only found in about 20% of plant species. Many nitrogenous compounds defend the plant through profound effects on the nervous system of animals, often interfering with nerve cell signaling in the central nervous system by binding to neurotransmitter receptors. Molecules such as nicotine, cocaine, heroin, and morphine are all alkaloids and act in this way. Other alkaloids such as those produced in the Nightshade family (Solanaceae) effect nerve cell signaling in the peripheral nervous system and can disrupt digestive processes of potential herbivores. In addition to the alkaloids, this class includes the cyanogenic glycosides. These potent deterrents release cyanide gas when the tissues containing the cyanogenic glycosides are ruptured.
Defense
How do plants defend themselves?
- Secondary Compounds: What kinds of molecules do plants make to defend themselves?
- Defense Overview: What are the kinds of defense pathways in plants?
- Defense Signaling: How does a plant match its response with a specific attacker?

